Transplant, revisited
A few months ago, I talked about the performance of calling Matlab from Python. Since then, I implemented a few optimizations that make working with Transplant a lot faster:
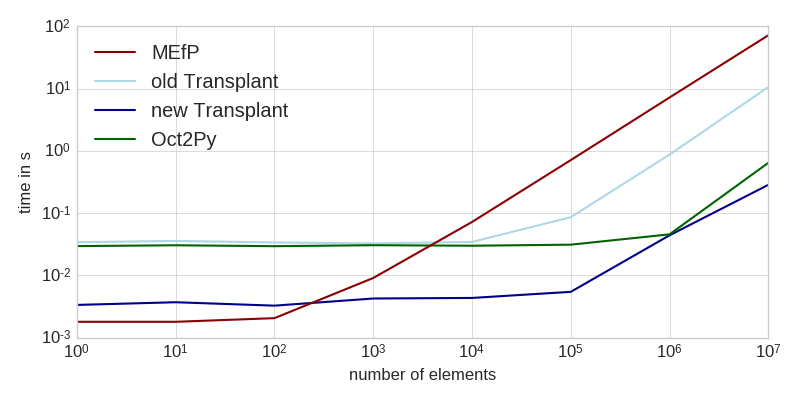
The workload consisted of generating a bunch of random numbers (not included in the times), and sending them to Matlab for computation. This task is entirely dominated by the time it takes to transfer the data to Matlab (see table at the end for intra-language benchmarks of the same task).
As you can see, the new Transplant is significantly faster for small workloads, and still a factor of two faster for larger amounts of data. It is now almost always a faster solution than the Matlab Engine for Python (MEfP) and Oct2Py. For very large datasets, Oct2Py might be preferable, though.
This improvement comes from three major changes: Matlab functions are now returned as callable objects instead of ad-hoc functions, Transplant now uses MsgPack instead of JSON, and loadlibrary instead of a Mex file to call into libzmq. All of these changes are entirely under the hood, though, and the public API remains unchanged.
The callable object thing is the big one for small workloads. The advantage is that the objects will only fetch documentation if __doc__ is actually asked for. As it turns out, running help('funcname') for every function call is kind of a big overhead.
Bigger workloads however are dominated by the time it takes Matlab to decode the data. String parsing is very slow in Matlab, which is a bad thing indeed if you're planning to read a couple hundred megabytes of JSON. Thus, I replaced JSON with MsgPack, which eliminates the parsing overhead almost entirely. JSON messaging is still available, though, if you pass msgformat='json' to the constructor. Edit: Additionally, binary data is no longer encoded as base64 strings, but passed directly through MsgPack. This yields about a ten-fold performance improvement, especially for larger data sets.
Lastly, I rewrote the ZeroMQ interaction to use loadlibrary instead of a Mex file. This has no impact on processing speed at all, but you don't have to worry about compiling that C code any more.
Oh, and Transplant now works on Windows!
Here is the above data again in tabular form:
| Task | New Transplant | Old Transplant | Oct2Py | MEfP | Matlab | Numpy | Octave |
|---|---|---|---|---|---|---|---|
| startup | 4.8 s | 5.8 s | 11 ms | 4.6 s | |||
| sum(randn(1,1)) | 3.36 ms | 34.2 ms | 29.6 ms | 1.8 ms | 9.6 μs | 1.8 μs | 6 μs |
| sum(randn(1,10)) | 3.71 ms | 35.8 ms | 30.5 ms | 1.8 ms | 1.8 μs | 1.8 μs | 9 μs |
| sum(randn(1,100)) | 3.27 ms | 33.9 ms | 29.5 ms | 2.06 ms | 2.2 μs | 1.8 μs | 9 μs |
| sum(randn(1,1000)) | 4.26 ms | 32.7 ms | 30.6 ms | 9.1 ms | 4.1 μs | 2.3 μs | 12 μs |
| sum(randn(1,1e4)) | 4.35 ms | 34.5 ms | 30 ms | 72.2 ms | 25 μs | 5.8 μs | 38 μs |
| sum(randn(1,1e5)) | 5.45 ms | 86.1 ms | 31.2 ms | 712 ms | 55 μs | 38.6 μs | 280 μs |
| sum(randn(1,1e6)) | 44.1 ms | 874 ms | 45.7 ms | 7.21 s | 430 μs | 355 μs | 2.2 ms |
| sum(randn(1,1e7)) | 285 ms | 10.6 s | 643 ms | 72 s | 3.5 ms | 5.04 ms | 22 ms |
